I am a Postdoc Researcher at the Eric and Wendy Schmidt Center, Broad Institute of MIT and Harvard, working with Prof. Caroline Uhler and Prof. Ramnik Xavier. My research aims to bridge machine learning and biology, focusing on developing computational methods to tackle unresolved biological problems. Actively contributing to the academic community, I serve as a reviewer for journals such as Genome Research, Advanced Science and PLOS Computational Biology, as well as top conferences such as ISMB and RECOMB.
Warning
Problem: The current name of your GitHub Pages repository ("Solution: Please consider renaming the repository to "
http://".
However, if the current repository name is intended, you can ignore this message by removing "{% include widgets/debug_repo_name.html %}" in index.html.
Action required
Problem: The current root path of this site is "baseurl ("_config.yml.
Solution: Please set the
baseurl in _config.yml to "Education
-
 Chinese Academy of SciencesPh.D. in System Science, Academy of Mathematics and Systems ScienceSep. 2017 - 2022
Chinese Academy of SciencesPh.D. in System Science, Academy of Mathematics and Systems ScienceSep. 2017 - 2022 -
 University of Science and Technology of ChinaB.E. in Automation, School of Information Science and TechnologySep. 2013 - Jul. 2017
University of Science and Technology of ChinaB.E. in Automation, School of Information Science and TechnologySep. 2013 - Jul. 2017
Experience
-
 Broad Institute of MIT and HarvardPostdoctoral Researcher, Eric and Wendy Schmidt CenterNov. 2022 - Present
Broad Institute of MIT and HarvardPostdoctoral Researcher, Eric and Wendy Schmidt CenterNov. 2022 - Present -
 University of Newcastle, AustraliaVisiting Student, Department of Electrical and Computer EngineeringJune 2016 - Sep. 2016
University of Newcastle, AustraliaVisiting Student, Department of Electrical and Computer EngineeringJune 2016 - Sep. 2016
News
Selected Publications (view all )
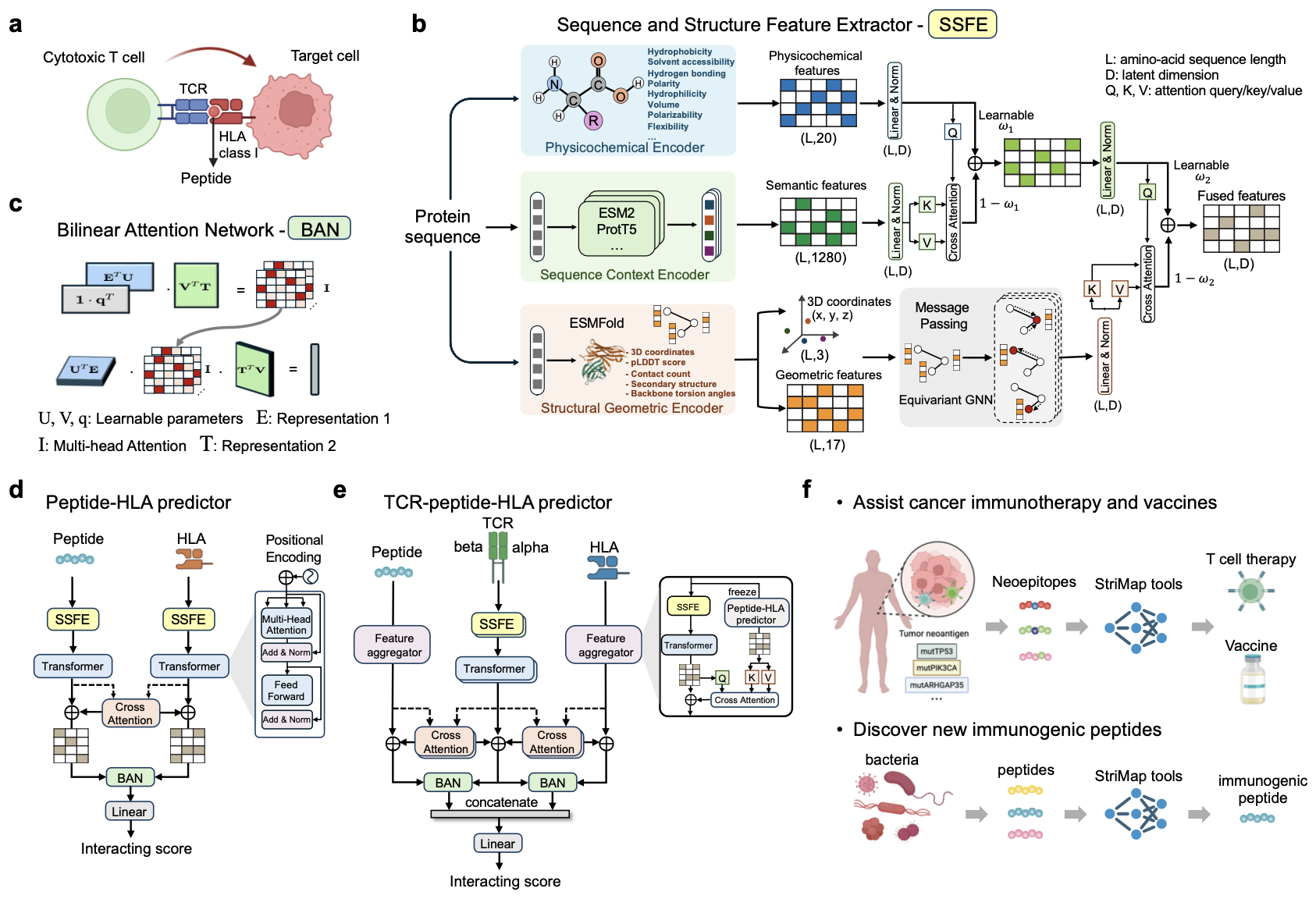
A structure-informed deep learning framework for modeling TCR-peptide-HLA interactions
Kai Cao, Rui Li, Martin Strazar, Eric M. Brown, Phuong N. U. Nguyen, Marie-Madlen Pust, Jihye Park, Daniel B. Graham, Orr Ashenberg, Caroline Uhler, Ramnik Xavier
bioRxiv 2026
A structure-informed deep learning framework for modeling TCR-peptide-HLA interactions
Kai Cao, Rui Li, Martin Strazar, Eric M. Brown, Phuong N. U. Nguyen, Marie-Madlen Pust, Jihye Park, Daniel B. Graham, Orr Ashenberg, Caroline Uhler, Ramnik Xavier
bioRxiv 2026
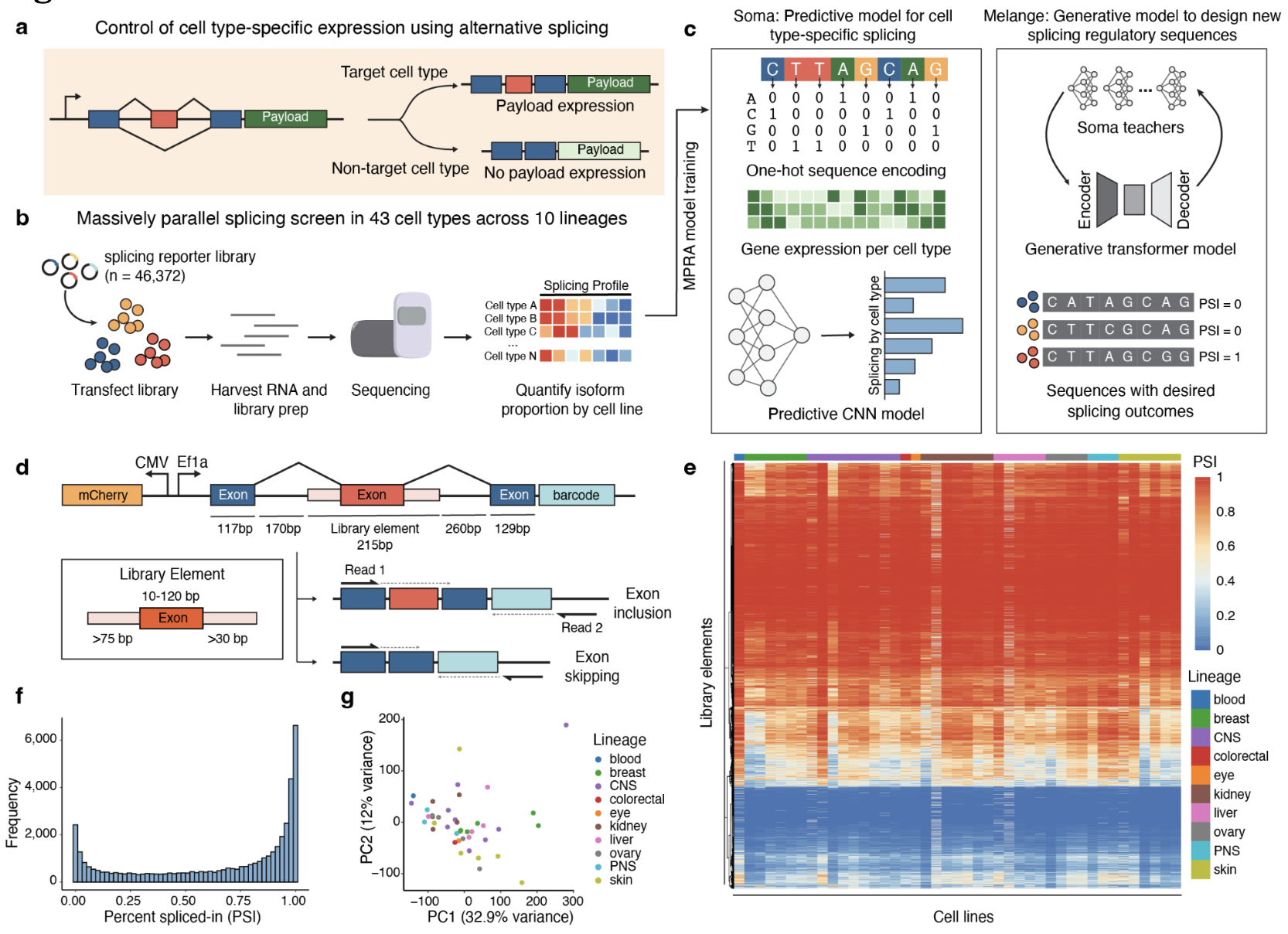
Generative Design of Cell Type-Specific RNA Splicing Elements for Programmable Gene Regulation
Xi Dawn Chen*, Maile Jim*, Mounica Vallurupalli*, Kai Cao*, Andrea Navarro Torres, Jing Wesley Leong, Yifan Zhang, David Wollensak, Qiyu Gong, Jing Sun, Mehdi Borji, Gail Schor, Sofia Mrowka, Margaret Hu, Anisha Laumas, Jennifer A. Roth, Todd Golub, Fei Chen (* equal contribution)
bioRxiv 2025
Generative Design of Cell Type-Specific RNA Splicing Elements for Programmable Gene Regulation
Xi Dawn Chen*, Maile Jim*, Mounica Vallurupalli*, Kai Cao*, Andrea Navarro Torres, Jing Wesley Leong, Yifan Zhang, David Wollensak, Qiyu Gong, Jing Sun, Mehdi Borji, Gail Schor, Sofia Mrowka, Margaret Hu, Anisha Laumas, Jennifer A. Roth, Todd Golub, Fei Chen (* equal contribution)
bioRxiv 2025
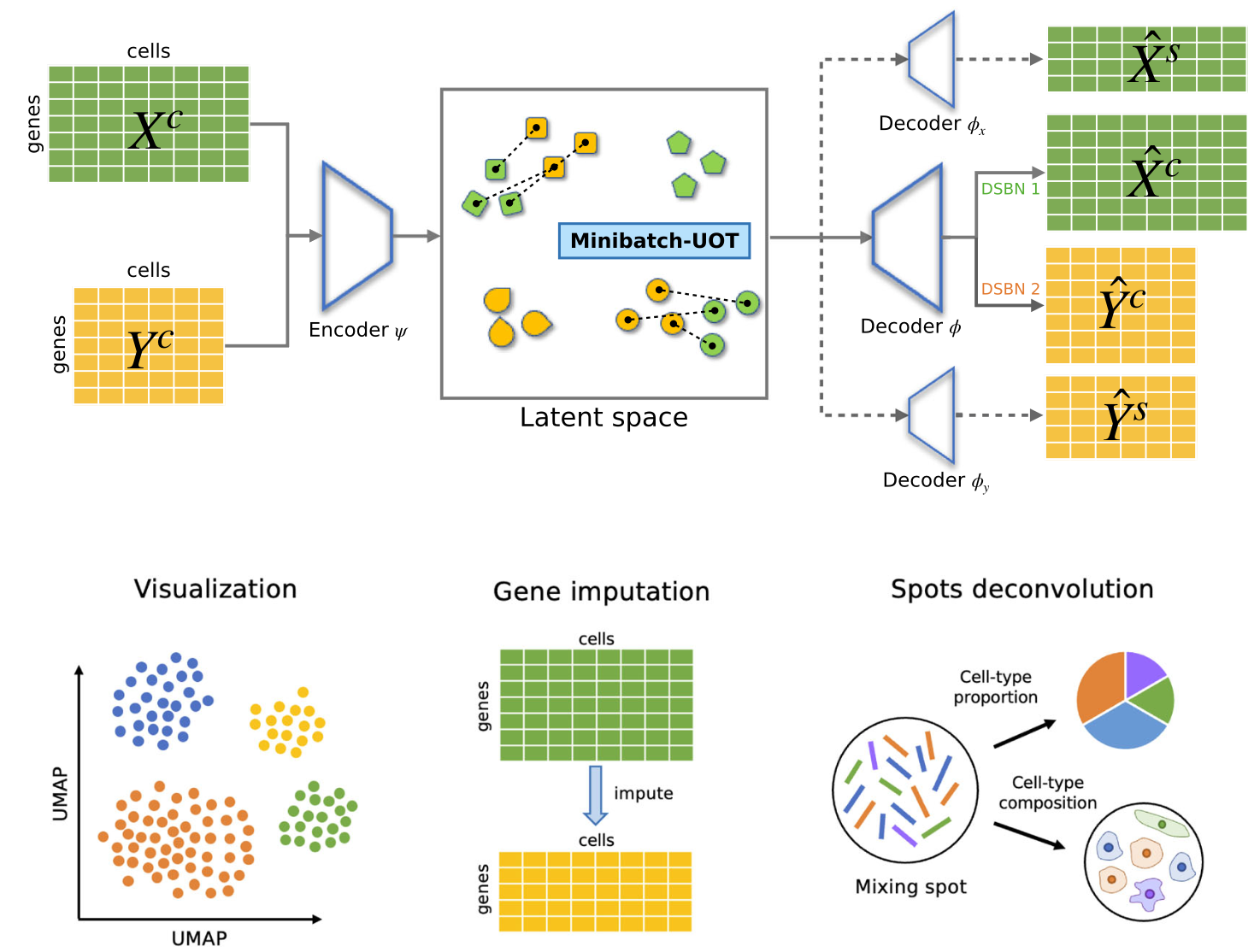
A unified framework for single-cell data integration with optimal transport
Kai Cao*, Qiyu Gong*, Yiguang Hong, Lin Wan (* equal contribution)
Nature Communications 2022 Editors’ Highlights
A unified framework for single-cell data integration with optimal transport
Kai Cao*, Qiyu Gong*, Yiguang Hong, Lin Wan (* equal contribution)
Nature Communications 2022 Editors’ Highlights
